New concepts in mass spectrometry measurements and interpretation
New concepts in mass spectrometry measurements and interpretation
Challenges for the structural interperation of ion mobility spectrometry data
Ion mobility spectrometry separates ions in the gas-phase according to their mobility in a buffer gas. The mobility depends on the balance between the acceleration by an electric field and the friction due to ion collisions with the buffer gas. Acceleration depends on the charge state (z) and friction depends on the ion size and shape. In low fields, the mobility value (K) can be related to a physical quantity called the collision cross section (CCS), which can be calculated theoretically for molecular models of the ions.
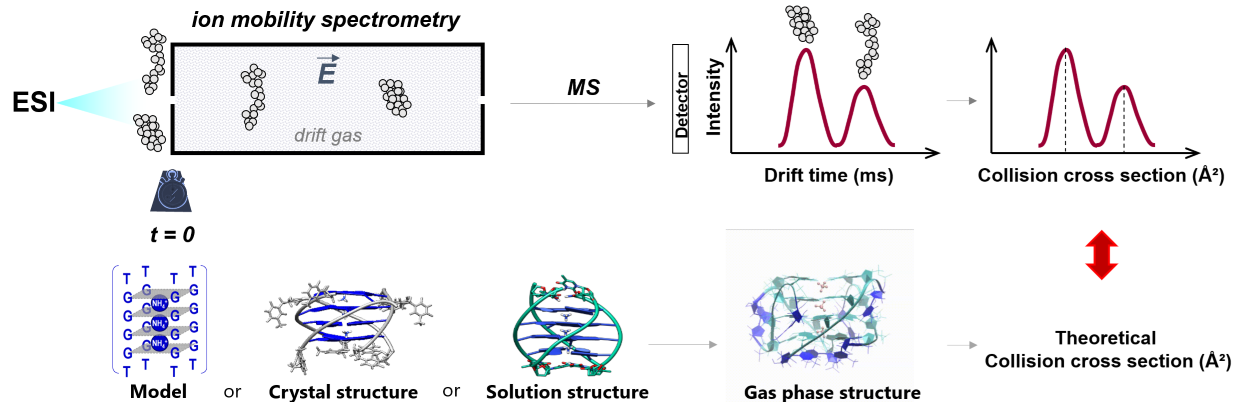
Previous work: We validated the parameterization to use for CCS calculations on oligonucleotides. We recommend (1) using mobility measurements and calculations in helium rather than in nitrogen and (2) calculating the theoretical CCS values either with EHSS or with TM using Siu’s re-parameterization [1]. We delineated how to reconstruct CCS distributions from the experimental arrival time distributions [2]. Finally, in a community paper, we provided recommendations to report ion mobility measurements, mobility and collision cross section values, for the various ion mobility measurement principles [3].
Current challenges:
- Understand how the electrospray process influences the charge states and conformation of the ions
- Delineate quantitative relationships between the gas-phase unfolding upon internal energy increase and the strength of the intramolecular forces
- Develop appropriate structural modeling approaches to obtain realistic gas-phase structures for CCS calculations
- Provide guidelines to sensibly interpret ESI-IM-MS measurements to infer what were the solution folding ensembles
References: [1] V. D’Atri, M. Porrini, F. Rosu, V. Gabelica, “Linking molecular models with ion mobility experiments. Illustration with a rigid nucleic acid structure”, J. Mass Spectrom. (2015), 50, 711. [2] Marchand A, Livet S, Rosu F, Gabelica V. “Drift Tube Ion Mobility: How to Reconstruct Collision Cross Section Distributions from Arrival Time Distributions?”, Anal. Chem. (2017) 89, 12674. [3] Gabelica V et al., “Recommendations for Reporting Ion Mobility Mass Spectrometry Measurements” Mass Spectrom. Rev. (2019), 38, 291.
Exotic fragmentation techniques (with electrons or lasers)
In mass spectrometry, pushing back the frontiers for the structural analysis of complex molecules and biomolecules requires one to increase the diversity of fragmentation and dissociation techniques. High mass resolution is necessary to distinguish the many fragments that a large molecule can produce, but even pushing the limits of mass resolution is useless if the proper fragments are not produced.
Project: Thanks to an R’Equip funding from the SNSF, by the end of 2024 we will acquire an instrument with ion activation network including higher energy collision-induced dissociation (HECID), slow-heating collision-induced dissociation (SHCID), electron capture dissociation (ECD), electron induced dissociation (EID) and ion activated dissociation (IAD). Two laser ports will enable us to couple the instrument with infrared and UV-vis tunable lasers for photo-dissociation (PD) or electronic ion spectroscopy. Multiple rounds of mass selection (MS/MS and MSn), combined with all the fragmentation techniques, will open the door to a myriad of possibilities for dissecting ion structures.
Novel ion spectroscopy approaches to measure chirality
Chirality is one of the most fascinating aspects of the molecules of Life. A molecule is chiral when its structure cannot be superimposed with its mirror image. Many biomolecules naturally occur only in one specific chiral form, called “enantiomer”. For example, DNA is chiral because it consists of chiral (D-) sugars, and its double helix turns towards the right, not the left. Proteins are chiral because they contain L-amino acids. In fact, many small molecule metabolites or natural products (secondary metabolites) are chiral in all kingdoms of Life. Consequently, when synthesizing compounds for applications in Life Sciences and Medicine, obtaining molecules with the desired chirality is of utmost importance. How to determine which chiral form (which enantiomer) is present in a sample?
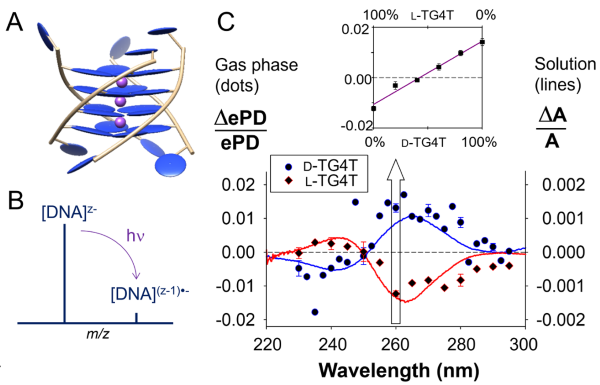
Previous work: In 2020, we showed that circularly polarized laser-induced electron photodetachment could generate electronic circular dichroism (CD) spectra of DNA structures in the gas phase [4]. This development of chiral action spectroscopy was the high-risk/high-gain work package of ERC CoG project “DNAFOLDIMS”. Although it was a breakthrough to show that this was even possible, admittedly the approach has one big limitation: using electron photodetachment as the single-photon action makes the approach applicable only to multiply charged anions, and thus only to relatively large molecules.
Project: We now aim to generalize chiroptical action spectroscopy to proteins, multiply charged cations, singly charged ions, or even neutrals.
Reference: [4] Daly S, Rosu F, Gabelica V, “Mass-resolved electronic circular dichroism ion spectroscopy”, Science (2020) 368, 1465.